
The Linux kernel has built-in packet filtering functionality through the Netfilter kernel subsystem. It can read and process packets by header information and filters the packet based on sets of programmable rules implemented by the firewall administrator.
JUPYTER NOTEBOOK ONLINE LEARN HOW TO CODE PC
Threats to Workstation and Home PC Security Using that, you can get a Launch Binder badge that you can add to your repository and launch it any time. Another example, this one R code, based on a gist shared on a twitter exchange can be seen here. That was done to produce this launchable repo after I forked one that had not been set up to use the Binder system initially.
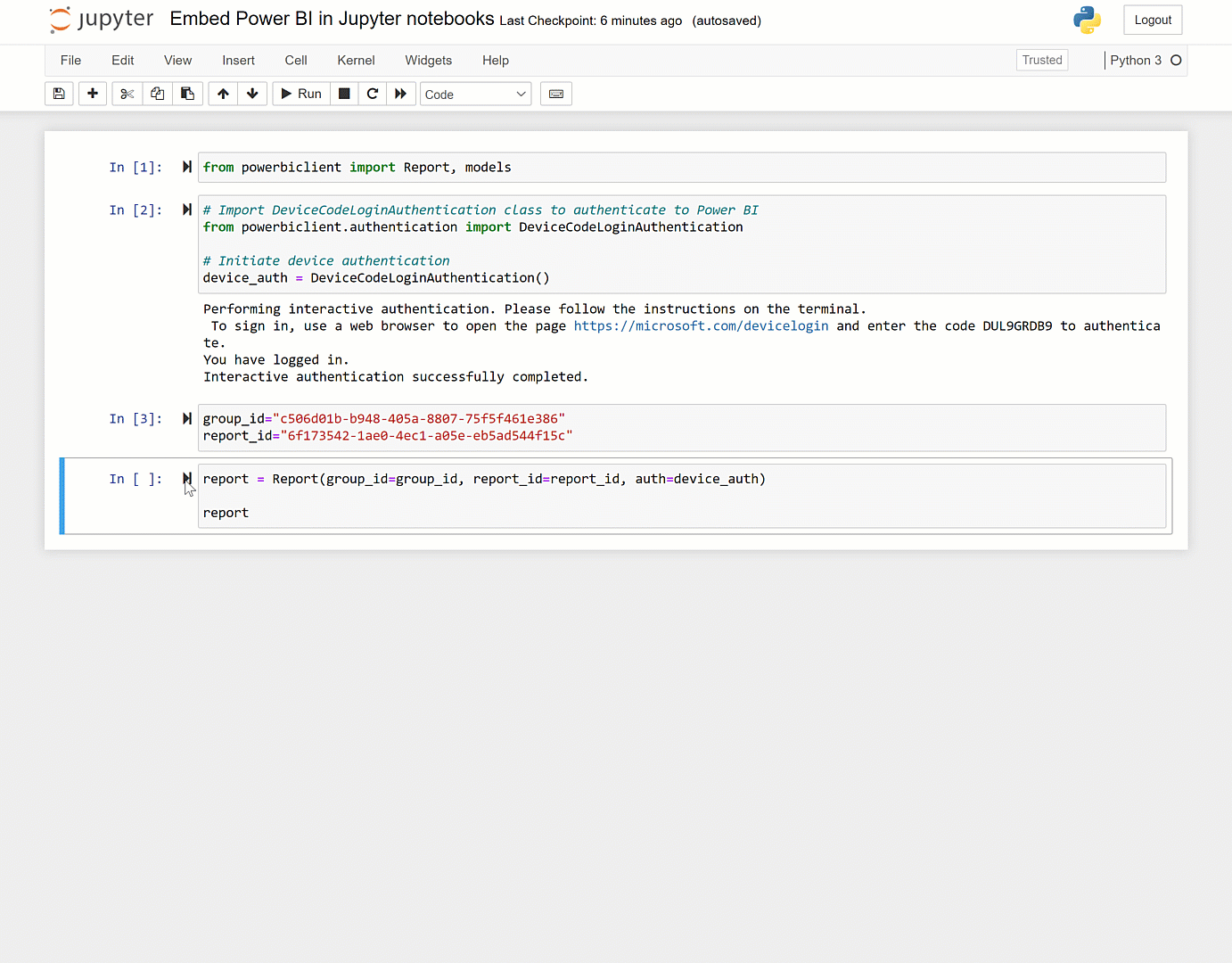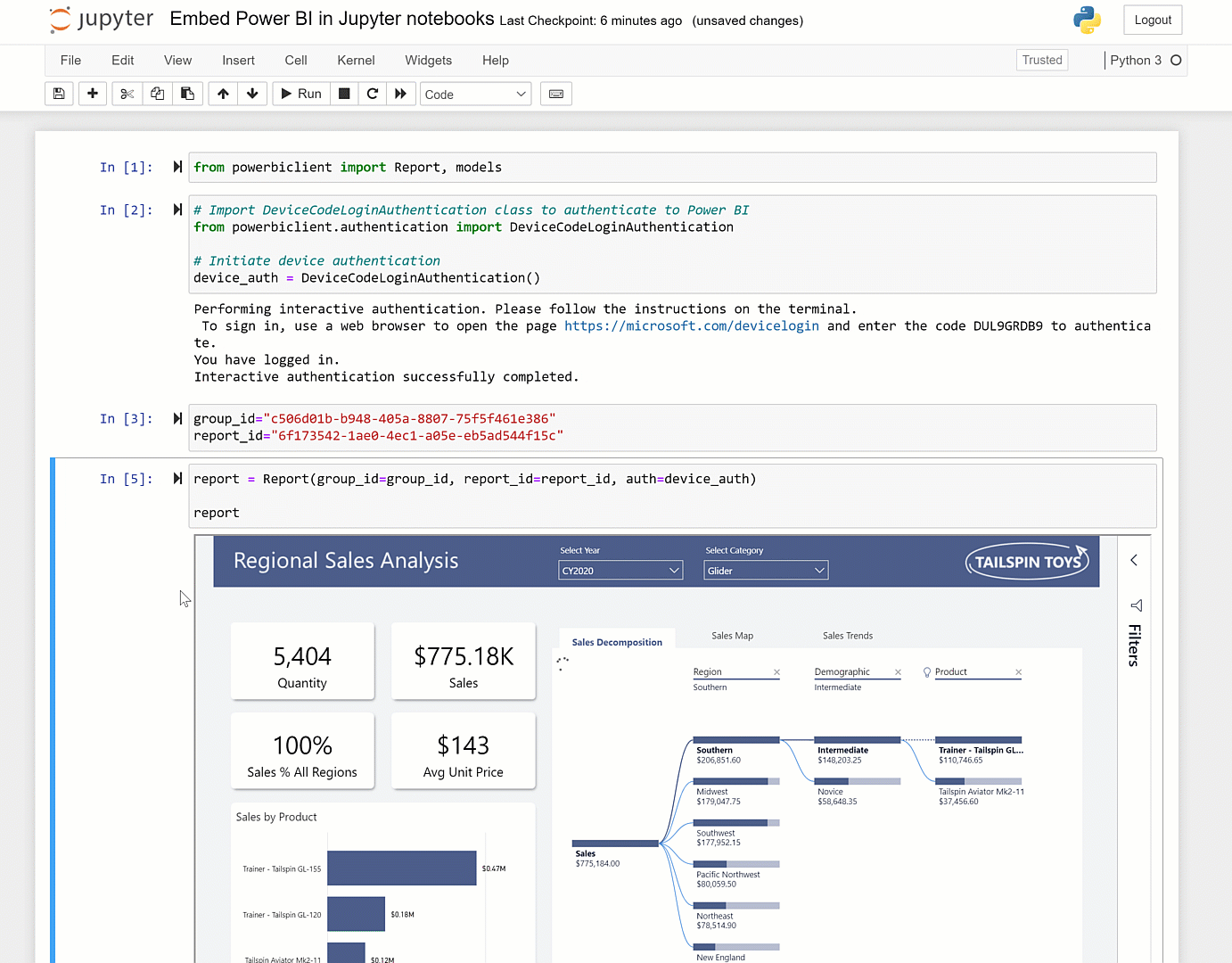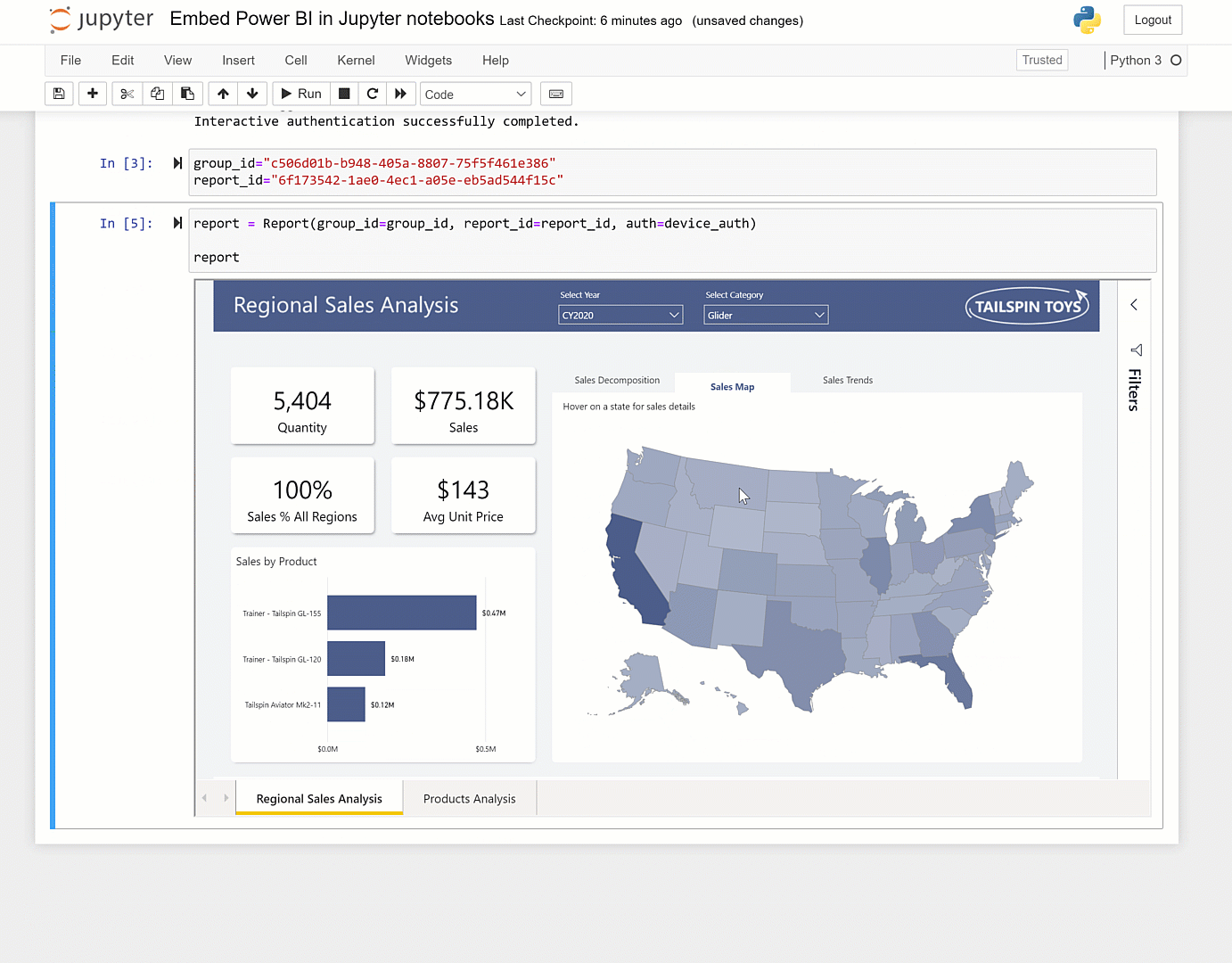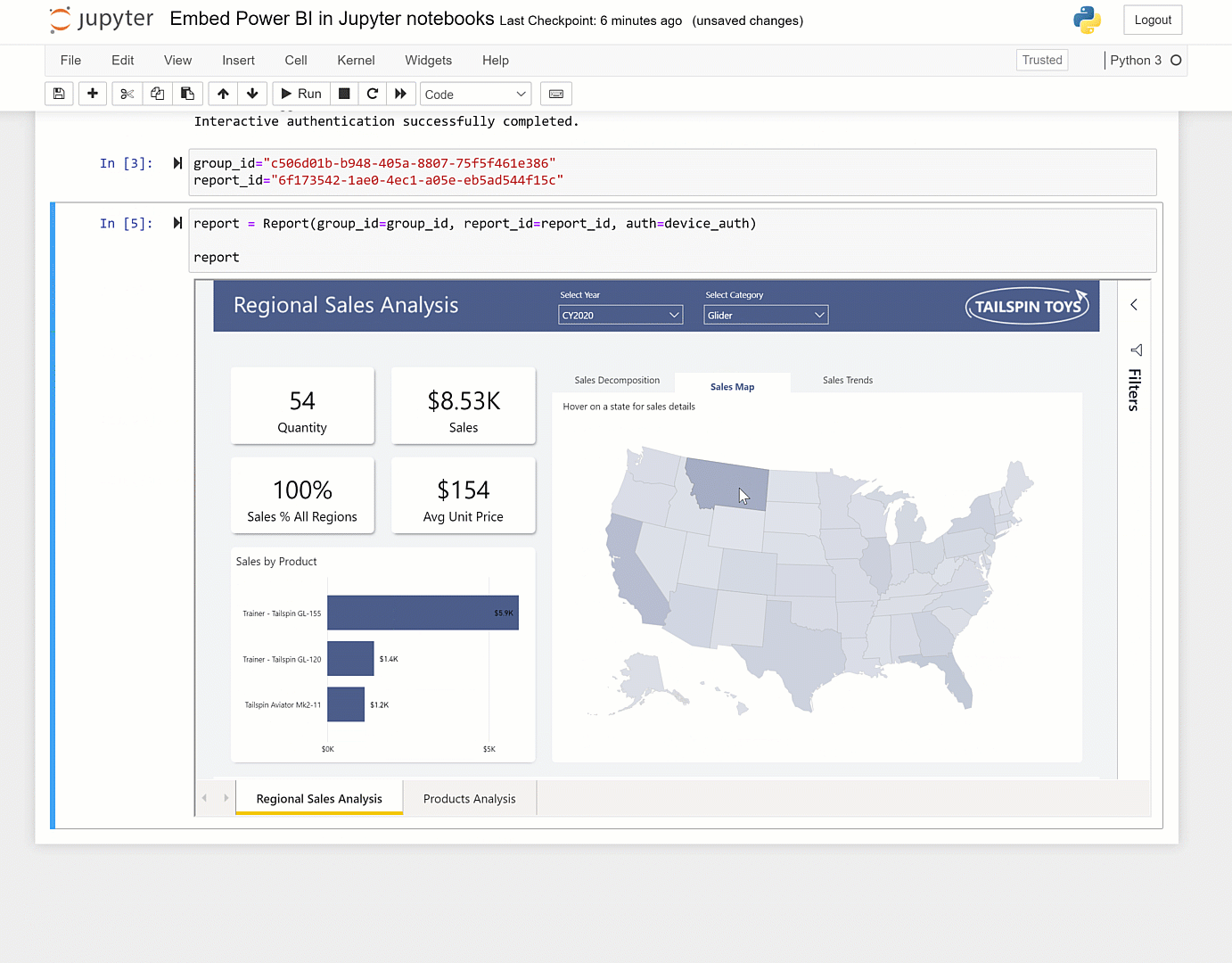
You can set it up to address that as guided by here and here. The caveat though is that unless it is fairly vanilla python in the notebook, you'll hit dependency issues. More information on the service can be found here, here, and here.Īt the page you can point the service at any Github repository. Or if you want others to be able to easily run that notebook.Ĭheck out highlighted in this Nature article here. If you want to run a notebook placed on GitHub: Using %run in the notebook provides a more full-featured Jupyter experience the use of %run to run a script in a Jupyter notebook is similar to the traditional way to run a script on the command line. Often you can run %run -help to get information on what arguments the script expects. Provide any necessary arguments for the script after its name.

Then you'd run the script in your Jupyter notebook with the following in a cell: %run fastq-to-fasta.py Similarly, if you want to bring the script in as a file you can call in the notebook using %run (or from the command line equivalent), use curl in the notebook cell and the script will be added to the current directory. More on other magic commands can be found here.
JUPYTER NOTEBOOK ONLINE LEARN HOW TO CODE CODE
More about using raw code via GitHub or Gists here and here.

That will pull in the code to the notebook's namespace when you execute it in a Jupyter notebook cell. Place what you extract from the address bar after %load to get something along the lines of this: %load You can get the URL for the raw code by browsing the script on GitHub or and pressing Raw in the toolbar just above the code. The trick with a Github or Gist-hosted script is to direct it at the URL for raw code. The source can either be a file on your computer or a URL. The IPython Magic command %load, as described in tip# 8 here, will replace the contents of the Jupyter notebook cell with an external script. If you just want to run Python code hosted on Github or in a Gist:
